by Sequentia
A solution to close the gap between data production and interpretation in transcriptomics. Obtain your full Differential Gene Expression and Gene Ontology report within a few hours.
Economic
perform your own RNA-Seq data analysis at a fraction of the average price
Scientifically robust and reproducible
uses the latest, peer-reviewed bioinformatics methods & algorithms
Responsive
run AIR from your mobile or tablet.
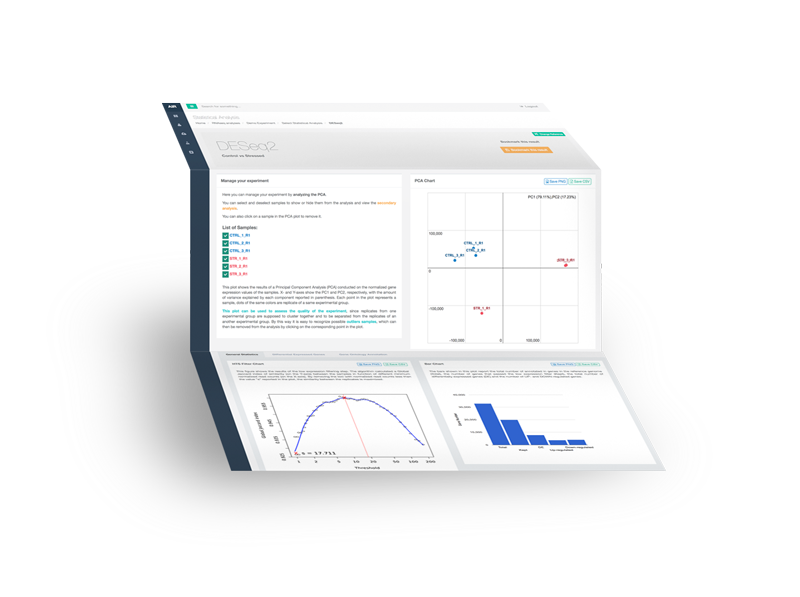
Full Functionality
navigate, download, save & share your results


What our users say
The software is user-friendly and the results are easy to see and export as publication-quality figures. A.I.R. is unique in its simplicity and accuracy. I anticipate that A.I.R. will become an indispensable everyday tool for RNA-Seq analyses."